Cell-free snythetic biology for combinatorial biosensor design
+++ positions filled +++ 12 Doctoral Candidate Positions for MSCA Doctoral Network
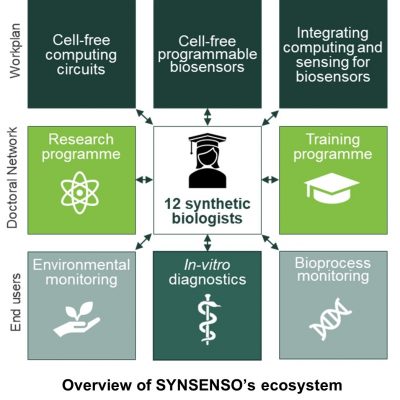
The Doctoral Network SYNSENSO “Cell-free synthetic biology for combinatorial biosensor design”, funded within the framework of the Marie Skłodowska-Curie Actions (MSCA), follows an interdisciplinary and cross-sectoral approach by bringing together academic and industrial experts from the fields of cell-free synthetic biology and molecular sensor design to develop novel combinatorial biosensors.
Five academic research groups and two industrial partners, coordinated by Technische Universität Darmstadt in Germany, join forces in SYNSENSO to create a mobility and training platform for young scientists by means of cross-site, interdisciplinary research projects. The DCs will work on individual research projects in order to devise new logic circuits in cell-free systems, build novel responsive elements for analyte detection and integrate logic circuits with response elements to build and test next-generation, combinatorial biosensors.
What we offer
The project offers the DCs research and training excellence in synthetic biology and biosensor design. The partners of SYNSENSO are leading research groups in synthetic biology and sensors design and their research institutes actively promote young researchers. The 6 academic research groups and 4 partners from the private sector join forces in SYNSENSO to create a platform of intersectoral and multidisciplinary mobility and training. To complement the academic and scientific goals of the DCs, the project offers customized research projects, structured interdisciplinary local and network-wide transferable skills training activities, and secondments at top-ranking European universities and industry partners.
Host: Prof. Heinz Koeppl, Technische Universität Darmstadt, Germany
Within this project energy-efficient RNA-based gates that can be cascaded will be designed and used as combinatorial sensors for long-noncoding RNAs.
Host: Prof. Tom de Greef, Eindhoven University of Technology, Netherlands
The goal of this project is to increase the sensitivity and specificity in the detection of analyte molecules through optimized ribozymes. Thereby neuronal networks will be used to enable faster and more cost-effective identification of suitable linker sequences for interesting analysis by reducing the need for time-consuming laboratory tests.
Host: Prof. Tom de Greef, Eindhoven University of Technology, Netherlands
The doctoral candidate will explore design principles for logic gates utilizing co-localization of regulatory RNA elements at the nanoscale. Within this project a cell-free platform based on integration of DNA origami nanostructures with E. coli will be developed.
Host: Prof. Heinz Koeppl, Technische Universität Darmstadt, Germany
This project is dedicated to taking cell-free lysates to the next level: overcoming the energy-limitation of cell-free lysates enabling more complex biosensors; understanding resource competition in bacterial production strains through expression of a regulator palette; minimize batch variability and maximum efficacy of regulators within the cell-lysate.
Host: Prof. Francesco Ricci, Tor Vergata Universita degli Studi di Roma, Italy
Within this project a cell-free biosensors for DNA repair enzymes will be developed Thereby the candidate will work on novel programmable DNA/RNA elements responsive to different DNA repair enzymes; cell-free transcription of different RNA sequences achieved with different DNA repair enzymes, orthogonal cell-free circuits responsive to DNA repair enzymes.
Host: Dr. Velia Siciliano, Istituto Italiano di Technologia, Naples, Italy
The doctoral candidate will evaluate new RNA-binding tools for translational control and for Boolean logic gates in cell-free systems. The goal is to apply the research results to a general-purpose platform for protease-based detection viral infection and to develop a low-cost paper-based biosensor for rapid infection detection.
Host: Prof. Francesco Ricci, Tor Vergata Universita degli Studi di Roma, Italy
This research project will explore novel DNA/RNA responsive elements with programmable and controllable input/output behaviour to reach a tuneable dynamic range by using different intrinsically disordered regions and eventually achieve digital-like response of DNA/RNA circuits with transient and reversible activation of responsive DNA/RNA elements.
Host: Dr. Ralf Strasser, Dynamic Biosensors GmbH, Munich, Germany (PhD awarded from TU Darmstadt)
Within this application-oriented project the doctoral candidate will evaluate cell-free expression systems for their use on biochips. The focus here is on improving assays for detection of conformational changes of RNA aptamers and the development of general method for attaching a recognition element to DNA using cell-free systems.
Host: Dr. Bruna Marini, Ulisse Biomed GmbH, Udine, Italy (PhD awarded from Tor Vergata Universita degli Studi di Roma)
Within this project a DNA/RNA responsive element with fluorescent detection for trastuzumab (mono-specific antibody), blinatumomab (bi-specific antibody) and EGFR will be developed. Thereby an orthogonal optical detection of all three targets in the same solution should be achieved.
Host: Prof. Karen Polizzi, Imperial College London, UK (will be funded through the UKRI Horizon Europe Guarantee Funding Scheme)
The work on this project includes the development of novel design principles for fast protein-based logic gates. Thereby an intein toolbox with well-characterized parts will be optimized for cell-free systems and the kinetics of intein splicing under different reaction conditions will be evaluated.
Host: Prof. Karen Polizzi, Imperial College London, UK (will be funded through the UKRI Horizon Europe Guarantee Funding Scheme)
The goal of the project is to establish new mechanism for harnessing the diversity of two-component signalling systems for biosensors in cell-free systems. A set of well-characterized parts for implementing phosphorylation cascades in cell-free systems should be developed and applied on a prototypical design of a fast biosensor for antimicrobial peptides.
+++ All positions have been filled. ++++
Disclaimer: Please note that due to a high volume of applications received, we are unable to contact each applicant individually regarding the status of their application.
